|
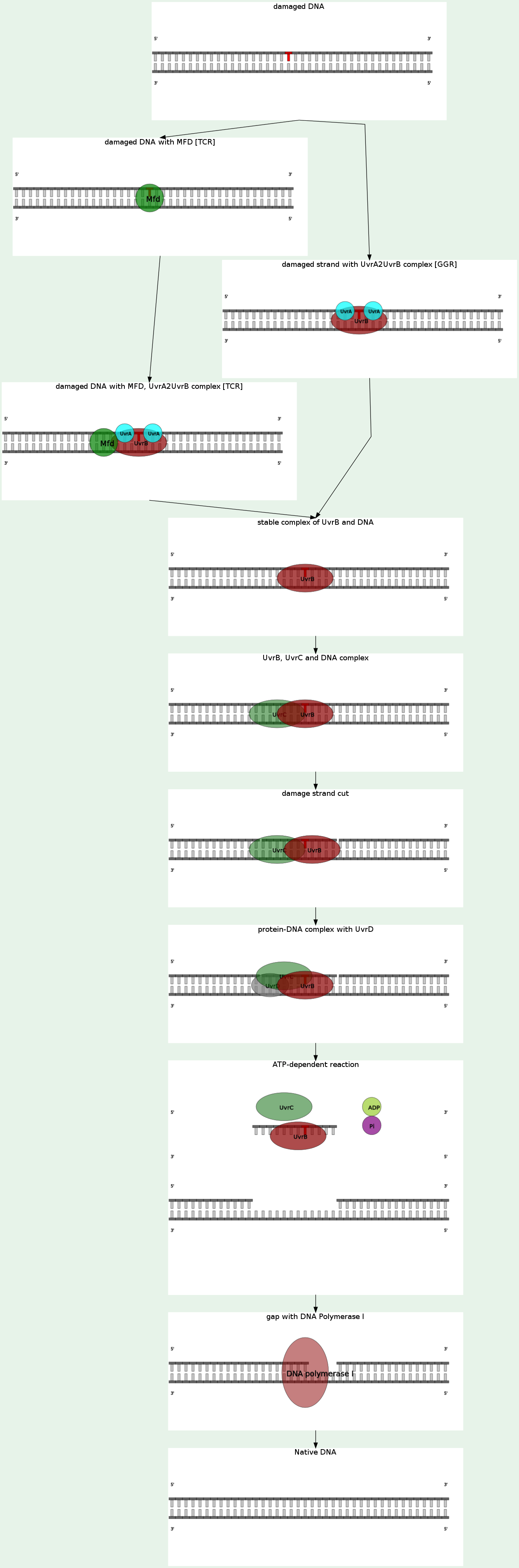
NER pathway removes UV-induced damages (mostly thymine dimers and 6-4-photoproducts) as well as other kind of DNA damages, which produse bulky distortions in the shape of DNA double helix. During the pathway a short single-stranded DNA fragment (containing a damage) is digested and rebuilt with use of undamaged strand as a template.
DNA constantly requires repair due to damage that can occur to bases from a vast variety of sources including chemicals but also ultraviolet (UV) light from the sun. Nucleotide excision repair (NER) is a particularly important mechanism by which the cell can prevent unwanted mutations by removing the vast majority of UV-induced DNA damage (mostly in the form of thymine dimers and 6-4-photoproducts). The importance of this repair mechanism is evidenced by the severe human diseases that result from in-born genetic mutations of NER proteins including Xeroderma pigmentosum and Cockayne's syndrome. While the base excision repair machinery can recognize specific lesions in the DNA it can correct only damaged bases that can be removed by a specific glycosylase, the nucleotide excision repair enzymes recognize bulky distortions in the shape of the DNA double helix. Recognition of these distortions leads to the removal of a short single-stranded DNA segment that includes the lesion, creating a single-strand gap in the DNA, which is subsequently filled in by DNA polymerase, which uses the undamaged strand as a template. NER can be divided into two subpathways (Global genomic NER and Transcription coupled NER) that differ only in their recognition of helix-distorting DNA damage.
Nucleotide excision repair has more complexity in eukaryotes. But the general principles upon which it operates are similar. There are 9 major proteins involved in NER in mammalian cells and their names come from the diseases associated with the deficiencies in those proteins. XPA, XPB, XPC, XPD, XPE, XPF, and XPG all derive from Xeroderma pigmentosum and CSA and CSB represent proteins linked to Cockayne syndrome. Additionally, the proteins ERCC1, RPA, Rad23, and others also participate in nucleotide excision repair.
Nucleotide excision repair can be categorized into two classes, global genome NER (GG-NER) and Transcription Coupled NER (TC-NER). Two different sets of proteins are involved in the distortion and recognition of the DNA damage in the two types of NER. In GG-NER, the XPC-Rad23B complex is responsible for distortion recognition, and DDB1 and DDB2 (XPE) can also recognize some types of damage caused by UV light. Additionally, XPA performs a function in damage recognition that is as yet poorly defined. In TC-NER, CS proteins CSA and CSB bind some types of DNA damage instead of XPC-Rad23B.
The subsequent steps in GG-NER and TC-NER are similar to each other and to those in NER in prokaryotes. XPB and XPD, which are subunits of transcription factor TFIIH have helicase activity and unwind the DNA at the sites of damages. XPG protein has a structure-specific endonuclease activity, which makes an incision 3’ to the damaged DNA. Subsequently XPF protein, which is associated with ERCC1 makes the 5' incision during the NER. The dual-incision leads to the removal of a ssDNA with a single strand gap of 25-30 nucleotides.
The resulting gap in DNA is filled by DNA polymerase δ or ε by copying the undamaged strand. Proliferating Cell Nuclear Antigen (PCNA) assists the DNA polymerase in the reaction, and Replication Protein A (RPA) protects the other DNA strand from degradation during NER. Finally, DNA ligase seals the nicks to finish NER.
|